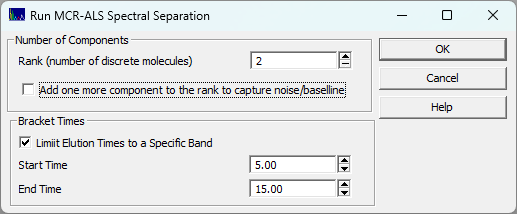
The dialog also offers an option to capture the noise and baseline. If the data have a high S/N and do not require baseline correction, this box should be unchecked.
PeakLab v2 Documentation Contents R2N Software Home R2N Software Support
Separate Compounds using MCR-ALS
The MCR-ALS algorithm is found in the ChromSpec | Separate Compounds using MCR-ALS.
Multivariate Curve Resolution (MCR-ALS)
MCR-ALS (Multivariate Curve Resolution - Alternating Least Squares) is an advanced chemometric technique used to mathematically unmix and resolve severely coeluting peaks in 3D spectrochromatographic data (DAD/PDA). Unlike standard non-linear peak fitting which models the geometric shape of a 2D slice, MCR-ALS is a purely data-driven matrix decomposition method that simultaneously resolves both the chromatographic profiles and the underlying pure spectra of overlapping compounds.
The Mathematical Foundation
At its core, MCR relies on the fundamental principle that the absorbance of a mixture is the linear sum of the absorbances of its pure components. The algorithm solves the classic bilinear matrix equation:
D = CS^T + E
Where:
D (Data Matrix): Your measured 3D dataset (Retention Time x Wavelength).
C (Concentration Matrix): The pure, resolved chromatographic elution profiles for each component.
S^T (Spectral Matrix): The pure, resolved spectra for each of those corresponding components.
E (Error Matrix): The unmodeled variance, consisting of baseline drift and instrumental noise.
How the "Alternating" Least Squares Works
The algorithm begins with an initial mathematical "guess" or seed (such as an estimated retention time for a hidden component). Because it is impossible to solve for both C and S simultaneously, the algorithm utilizes an alternating loop:
Assuming the Spectra (S) are correct, it performs a least-squares regression to calculate the Concentrations (C).
Assuming those new Concentrations (C) are correct, it performs a least-squares regression to calculate the Spectra (S).
This alternating seesaw continues iteratively until the error matrix (E) is minimized and the system converges on a stable solution.
Overcoming Rotational Ambiguity in PeakLab
A primary weakness of raw MCR is "rotational ambiguity"—the fact that a single matrix can have an infinite number of mathematically valid, but chemically impossible, solutions (such as predicting negative concentrations). To force the algorithm to find the true chemical reality, the PeakLab MCR-ALS implementation relies on three critical guardrails:
1. Non-Negative Least Squares (NNLS) Constraints:
PeakLab enforces strict non-negativity on both the Concentration (C) and Spectral (S) matrices during every iteration. This guarantees that the algorithm will never attempt to resolve a compound with a negative absorbance or a negative concentration.
2. Dynamic Gaussian Seeding:
Rather than relying on random noise or the purest variables to start the calculation, PeakLab intelligently seeds the initial concentration matrix using targeted Gaussian estimations. This gives the optimizer a massive head start and prevents the algorithm from collapsing into local mathematical minima.
3. Targeted Time Windowing:
Because MCR-ALS looks at the global variance of the entire matrix, distant peaks and shifting baselines can easily confuse the optimizer. PeakLab allows you to specify strict Start and End time cutoffs. By isolating a specific coeluting cluster, the algorithm is forced to focus its entire resolving power exclusively on the local overlap.
4. The Normalization Constraint (Solving Scale Ambiguity)
Because MCR-ALS factors the experimental data into two separate components (Concentration and Spectra),
it suffers from "Scale Ambiguity." Without constraints, the algorithm can arbitrarily inflate
the Y-axis of the chromatogram while proportionally shrinking the Y-axis of the spectrum, all while maintaining
a mathematically perfect fit. To ensure your resolved peaks have quantitative meaning, PeakLab automatically
applies a Normalization constraint to the Spectral matrix. This locks the maximum absorbance of every
resolved spectrum to exactly 1.0. Because the spectral scale is fixed, the resolved chromatograms are
forced to carry the exact quantitative magnitude (Absorbance Units) and area of your original experimental
data.

The dialog also offers an option to capture the noise and baseline. If the data have a high S/N and
do not require baseline correction, this box should be unchecked.
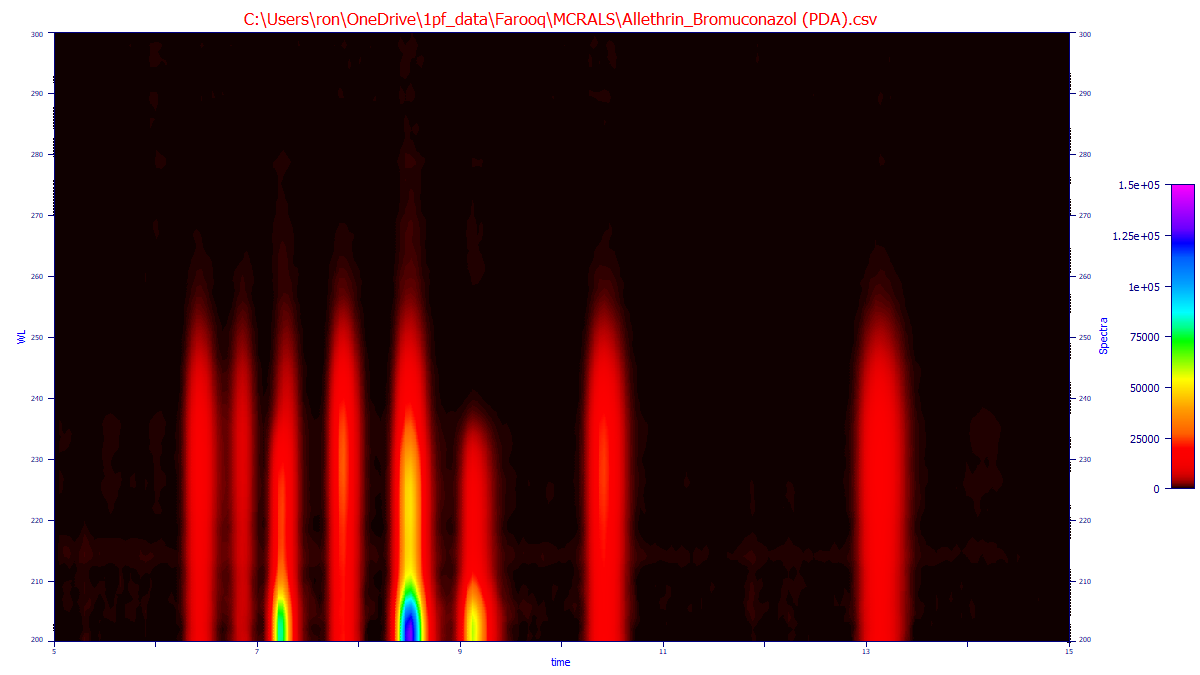
Example
This is the 3D data of a mix of allethrin and bromuconazol. One of these compounds have four stereoisomers, and the other has eight, and as such a rank of 2 (two compounds) is appropriate.
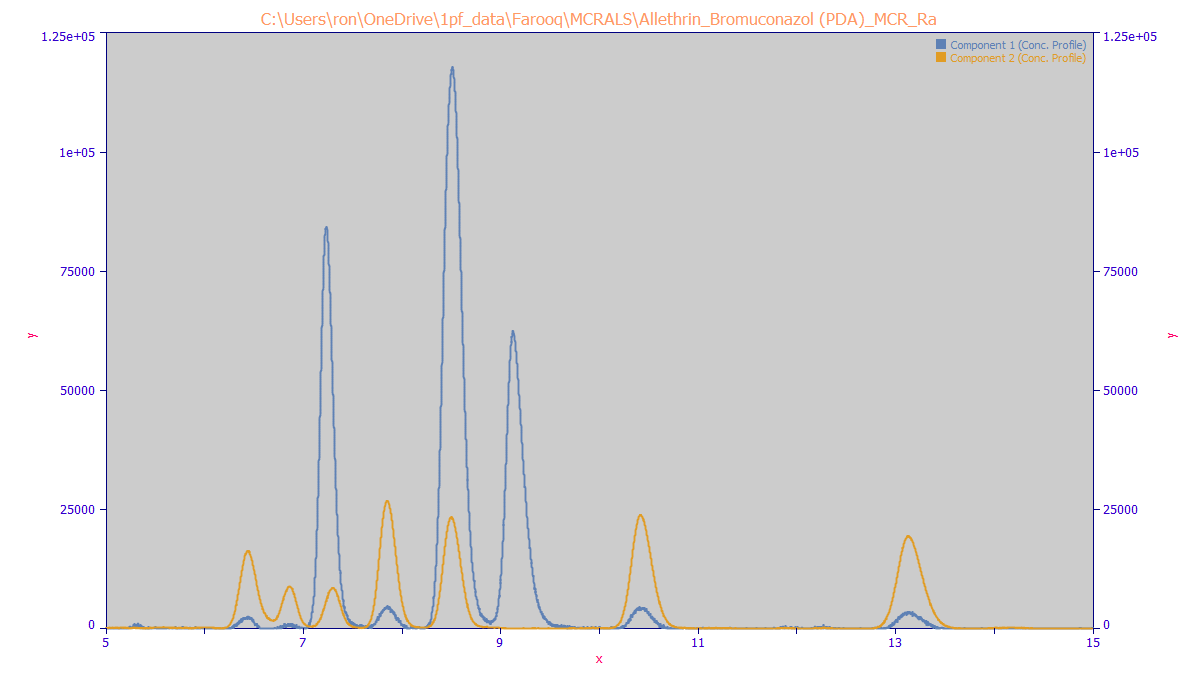
The MCR-ALS algorithm produces the following chromatographic separation of the two components:
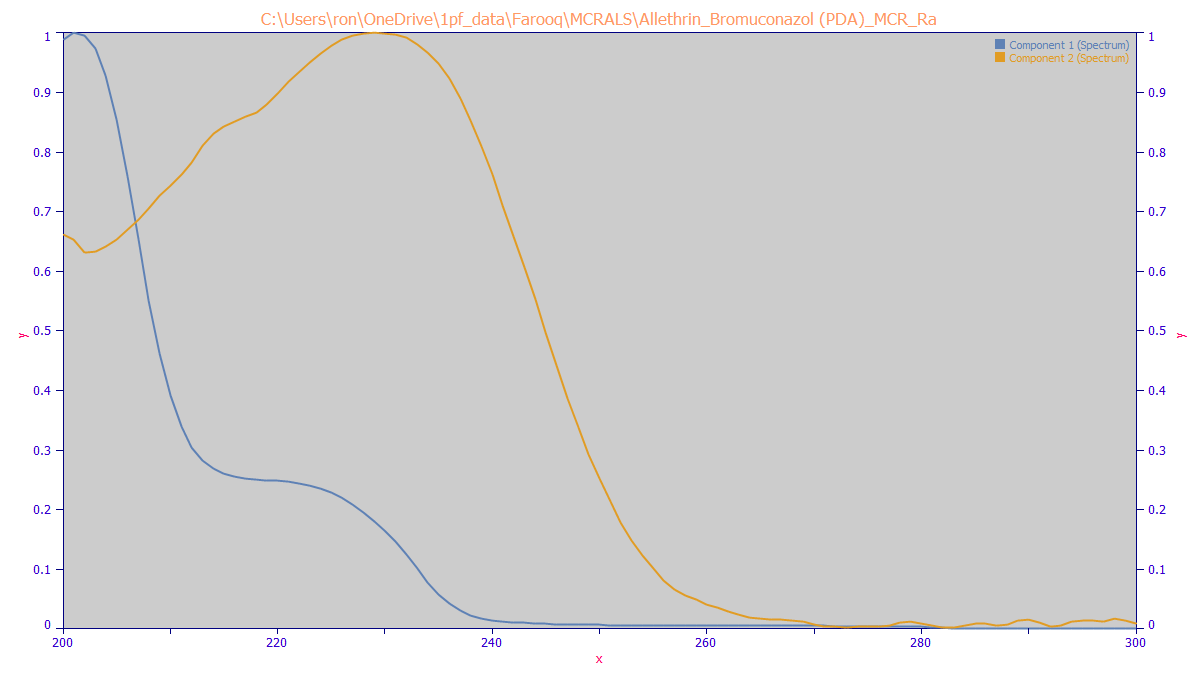
The MCR-ALS algorithm produces the following spectroscopic separation of the two components:
Acknowledgements and Thank You
The PeakLab authors would like to thank Dr. M. Farooq Wahab, Dr. Daniel W. Armstrong, and Siddharth Jaya Sajeevan J of the University of Texas, Arlington, for sharing this data from their paper. Their references are below:
Siddharth Jaya Sajeevan J, M. Farooq Wahab, *Daniel W. Armstrong, Selective Chemometric Elimination of
Co-Eluting Components in Chiral and Achiral Liquid Chromatographic Analyses, Analytical Chemistry, Vol
97/Issue 34, August 21, 2025
https://pubs.acs.org/doi/abs/10.1021/acs.analchem.5c03634
M. Farooq Wahab, Garrett Hellinghausen, Daniel W. Armstrong, LC GC, Progress in Peak Processing, May 01,
2019, Volume 32, Issue 5, pg 22–28
https://www.researchgate.net/profile/Garrett-Hellinghausen/publication/333639563_Progress_in_Peak_Processing/links/5cf871cc299bf1fb185bb7ab/Progress-in-Peak-Processing.pdf