PeakLab v2 Documentation Contents R2N Software Home R2N Software Support
MPPF (Massively Parallel Peak Fitting)
PeakLab’s Massively Parallel Peak Fitting (MPPF) engine is designed to handle the massive computational weight of fitting natural materials, proteins, peptides, and other highly complex samples. By utilizing the main direct fitting strategy (Fit with Full Data), HDPF allows you to simultaneously fit up to 1,000 discrete , overlapping, and hidden peaks in a single dataset.
Because each peak can require up to five independently fitted parameters, a 1,000-peak model requires the algorithm to construct and invert a 5,000 x 5,000 dense matrix during every single iteration. To solve mathematical problems of this magnitude at practical speeds, PeakLab employs a multi-tiered parallel processing architecture.
Parallel Processing
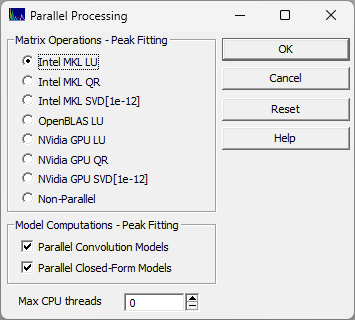
These parallel options are configured in the Edit menu's Parallel Processing option.

The Two Tiers of Parallelization
PeakLab achieves high performance by offering parallel computation at three different levels simultaneously. However, because your CPU has a finite number of physical cores, these levels can sometimes compete for resources. Understanding this hierarchy is the key to tuning your fitting speeds:
1. Dataset Parallelism (Each Data Set): PeakLab can independently fit up to 25 separate data sets on completely separate CPU threads at the same time.
2. Matrix Parallelism (Each Iteration in Each Data Set): The most intense calculation—the actual inversion of the dense matrix—is distributed across multiple cores or offloaded to a GPU.
3. Algorithmic Parallelism (Each Iteration in Each Data Set): Within a single dataset, the computation of the peak functions and their partial derivatives at each data point can be parallelized.
Configuring Matrix Operations (The Inverses)
The Matrix Operations section specifies exactly how the MPPF engine handles the massive matrix inversions.
LU Decomposition (Maximum Speed):
LU is the standard workhorse for matrix inversion and generally provides the fastest performance for well-behaved data. PeakLab offers both the Intel MKL LU and the OpenBLAS LU. Depending on your specific processor architecture (for example, the latest generation Intel processors), you may find that the OpenBLAS LU slightly outperforms the native Intel MKL.
QR Decomposition (Maximum Stability):
While QR decomposition is computationally heavier and slower than LU in single-threaded environments, it scales exceptionally well across multiple cores. It offers superior numerical stability for highly complex, dense fits.
SVD [1e-12] (The Safety Net):
Singular Value Decomposition (SVD) is the most computationally expensive option, but it is mathematically stable. You should switch to SVD if your 5,000 x 5,000 matrix becomes ill-conditioned (which can occur when hundreds of peaks are being fitted and there are coelutions) and the LU or QR solvers fail.
CPU vs. NVIDIA GPU:
If your workstation is equipped with a compatible NVIDIA graphics card, you can offload these massive matrix inversions entirely to the GPU, freeing up your CPU to handle the algorithmic functions and multi-dataset scheduling. Please note that PeakLab does not ship with the NVIDIA CUDA DLLs. You must independently install the NVIDIA development platform on your machine.
Configuring Model Computations (Function/Derivatives, IRFs)
This section controls the algorithmic parallelization—calculating the mathematical values of the peaks themselves.
Parallel Convolution Models:
If your models contain an Instrument Response Function (IRF), you will probably want to have this parallel option enabled. Deconvolving an IRF requires every single peak to undergo an independent Fourier convolution. This is computationally intensive. Parallelizing this specific step almost always yields significant performance gains, even when fitting may data sets dimultaneously. The internal thread scheduler is designed to keep the workload manageable.
Parallel Closed-Form Models:
When fitting a single, massive dataset, this may speed up the calculation of closed-form peak models. if you are simultaneously fitting many datasets at once (Top Tier parallelism), you will probably want to turn this option off. Otherwise, competing threads may slow your CPU.
Performance Tuning Examples
One Data Set
N = 8K, n peaks = 165, n parameters = 649, Fit with Full Data
Model Inverse NonConv Conv MatrSize Iters
Fstat Time(s)
GenHVL Intel MKL LU Yes N/A 649 x 649 500 14264 23.82
GenHVL Intel MKL LU No N/A 649 x 649 500 14264 18.71
GenHVL Intel MKL QR No N/A 649 x 649 500 14264 21.93
GenHVL Intel MKL SVD No N/A 649 x 649 500 12797 164.65
GenHVL OpenBLAS LU No N/A 649 x 649 500 14264 18.07
GenHVL Single-Threaded No N/A 649 x 649 500 14264 93.67
Twelve Data Sets
Each N = 8K, n peaks = 165, n parameters = 649
Model Inverse NonConv Conv MatrSize Iters
Fstat Time(s)
GenHVL Intel MKL LU No N/A 649 x 649 500 14264 54.0
GenHVL Intel MKL QR No N/A 649 x 649 500 14264 135.0
GenHVL Intel MKL SVD No N/A 649 x 649 500 12797 1590.5
GenHVL OpenBLAS LU No N/A 649 x 649 500 14264 53.5
GenHVL Single-Threaded No N/A 649 x 649 500 14264 225.0
Observations
The LU algorithm, either the Intel MKL or the OpenBLAS offers exceptional performance. The Intel MKL LU is the PeakLab default.
The QR algorithm has little additional cost with just one data set, but some of the performance is lost when many data sets are being fitted. The assumption is that the more computationally intensive QR is using a higher percentange of an iteration's duration, and negating some of the benefit of the multiple data set parallelization.
The SVD algorithm would be expected to use almost all of an iteration's duration. That is seen
with the much higher times on both the one and twelve data set fits. There is still a benefit from the
multiple data set parallelization, but it is small.
The single threaded matrix inverse is expensive for one data set, far less so for twelve identical
sets. In the twelve data set case, one realizes an indirect multithreading during the inverse computations
as many single threaded inverses will be computed simultaneously.
Unless you are using a math-intensive peak model, the cost of multithreading Parallel Closed-Form
Models is higher than any gain in performance, even when just one data set is being fitted.