PeakLab v2 Documentation Contents R2N Software Home R2N Software Support
Extract, Consolidate, and Fit High Resolution LC-MS MS1 Data (Qualitative)
Qualitative Feature-Based XIC Extraction (OpenMS)
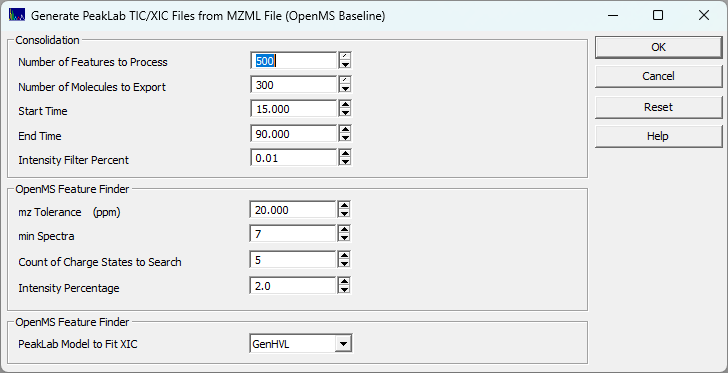
This is the MassSpec | Extract and Fit Qualitative HRMS XICs (OpenMS)... option. It utilizes the OpenMS FeatureFinderAlgorithmPicked to extract Extracted Ion Chromatograms (XICs) from the raw mass spectrometry data. You must first import MS data using the MassSpec | Import MZML data... or the MassSpec | Import Agilent data.ms Data... option.
Important Caveat: Qualitative Reference vs. Quantitative Analysis
PeakLab offers this native OpenMS feature finder primarily as a reference to demonstrate the current industry standard for template-based isotopic and charge-state feature finding.
Because OpenMS relies on a "top-down" 2D isotopic template match rather than PeakLab’s advanced "bottom-up" 2D shape-correlation (the IRD algorithm) standard feature finders often fragment broad, asymmetric, or overloaded signals into redundant, overlapping features. When these features are summed, it results in an "additive overshoot" that physically exceeds the raw Total Ion Chromatogram (TIC) data—meaning the exact same raw electrons are being double-counted.
Consequently, while this OpenMS baseline is a powerful tool for qualitative feature identification, it is not mathematically constrained to the conservation of mass. For high-fidelity quantitative work, we strongly recommend using the PeakLab IRD algorithm.

Consolidation (Limits & Filtering)
Note: In the context of the OpenMS procedure, "Consolidation" refers to the sorting and capping of final features to prevent memory exhaustion, rather than the mathematical clustering of raw strands seen in the IRD pipeline.
Number of Features to Process
The maximum number of raw features the OpenMS engine is allowed to evaluate.
Number of Molecules to Export
Caps the final list of exported XICs to the top N features (ranked by total area). Lowering this number is the most effective way to strip out the redundant "echo" fragments generated by the OpenMS algorithm on overloaded peaks.
Start Time / End Time
Restricts the analysis to a specific retention time window. Set both to 0.000 to process the entire time range in the separation.
Intensity Filter Percent
The primary noise floor, defined as a percentage of the global maximum intensity. To prevent the OpenMS engine from attempting to fit the absolute noise floor (which causes combinatorial memory failure), PeakLab enforces a minimum of 0.005%.
OpenMS Feature Finder (Algorithm Settings)
These settings are passed directly to the OpenMS engine to constrain its template matching.
mz Tolerance (ppm)
The mass extraction window. PeakLab automatically converts this parts-per-million value into the Dalton window required by the underlying OpenMS C++ library.
min Spectra
The absolute minimum number of unbroken, consecutive scans required to validate a feature. Raising this value (e.g., from 3 to 7 or 10) forces OpenMS to be stricter, which can help prevent it from spawning false features on the noisy tails of massive Orbitrap peaks.
Count of Charge States to Search
The upper bound of the charge states the isotopic template will attempt to match.
Intensity Percentage
The relative intensity threshold for including isotopic peaks within the Averagine model template.
Fitting
PeakLab Model to Fit XIC
Selects the non-linear mathematical model used to fit the extracted OpenMS features. The GenHVL (Generalized Haarhoff-Van der Linde) natively handles the fronting and tailing characteristic of concentration-impacted LC peaks.
Diagnostic Output (The "Black Box" Distinction)
If you have specified a PeakLab Export path, this procedure will output high-resolution diagnostic PNGs and an Excel-compatible CSV summarizing the OpenMS features.
Please see the IRD Algorithm help for a description and examples of these diagnostic outputs.
The Missing Unconsolidated Data
Users familiar with the PeakLab IRD algorithm will notice the absence of an option for the unconsolidated_strands plot and CSV. This is a fundamental architectural difference. OpenMS is a "one-step" process: it goes straight from the raw mass/time grid to a finished feature using a 2D bounding box. It never extracts individual isotopic strands first. Because OpenMS acts as a "black box," there is no mathematical way to verify how the underlying isotopic traces were grouped. The diagnostic files generated here represent the maximum amount of transparency the standard OpenMS library is physically capable of providing.