PeakLab v2 Documentation Contents R2N Software Home R2N Software Support
Filter DAD/PDA Baseline
The Filter DAD/PDA Baseline... in the ChromSpec menu is used to to baseline correct all of the chromatograms in a DAD/PDA matrix. This option is also presented in each import procedure. You must first import spectra for this procedure to be available.
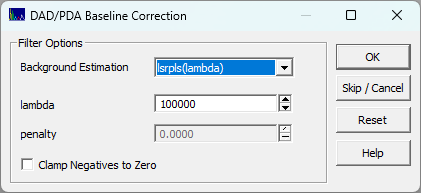
In this option you will be presented with a small background correction dialog:

This will use a Whittaker
baseline method of your choice to baseline correct all of the chromatographic spectra in the file.
There will be one baseline correction computed for every wavelength in the file. This may perform many
hundreds of automated baseline corrections. This is essential if you are working with 3D chromatographic
spectra from gradient experiments where there is a baseline that strongly varies with the mobile phase
strength.
Please note that there may be substantial differences in baselines at the different wavelengths, and
while the Whittaker lambda is generally a function of N, the number of data points, lambda proportional
to the log(N), it will also vary with the width of the peaks.
For this reason we suggest one of the adaptive weighting Whittaker procedures. They are easy to spot
since there is only the single lambda parameter. For chromatographic PDA spectra, arpls, brpls, lsrpls
are often simplest and most useful because of this adaptive function. These adaptive weights will mean
differences in how the baseline is estimated across the different wavelengths, however, and if it is important
to treat all wavelengths identically, you should use an algorithm with a second term that will control
how high or low the baseline will sit in the noise of the data, such as psalsa.
Manual Determination of Baseline Parameters
You can use the Skip/Cancel to omit this step and then use the Import Chromatographic Data Sets... option at wavelengths of interest to create data sets that can be individually evaluated with the Baseline option. There is a baseline optimization method where you can manually specify an optimal human designed baseline and a genetic algorithm optimization will be made to determine the lambda and penalty values that most closely reproduce the human baseline.
Graphical Determination of Background Correction
The SymLog (the third item in clicking the LogZ sequence in the View DAD/PDA 3D Matrix option
![]() The Logarithmic Z in the View DAD/PDA 3D Matrix plots offers a third SymLog Z-scale option.
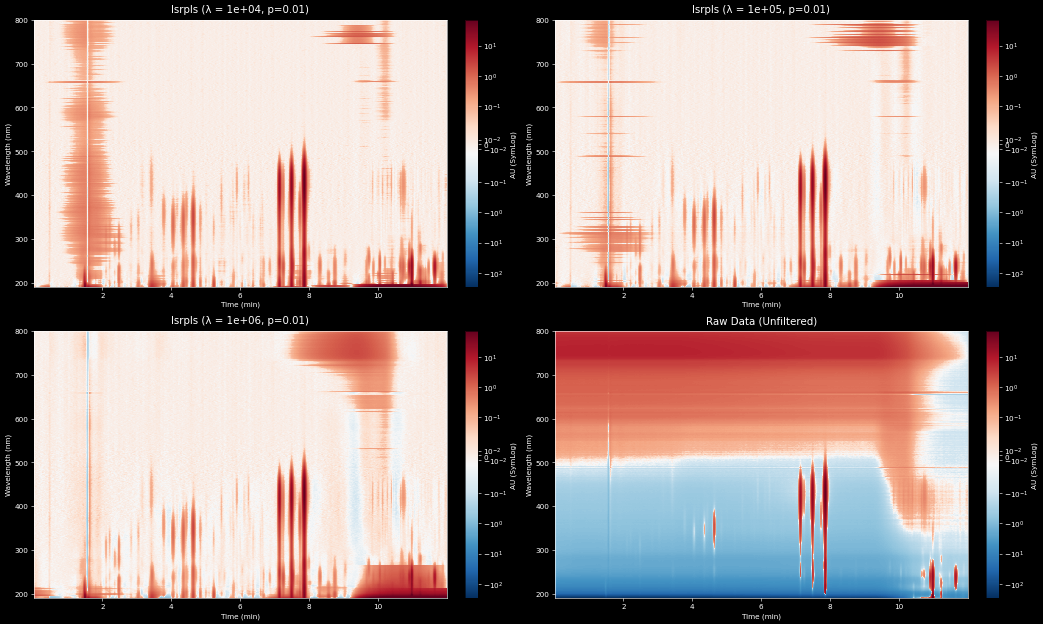
With the RdBu_r colormap, or another that goes to white at 0, you can judge the efficacy of the background
subtraction. Here we see the efficacy of the lsrpls at three different lambdas and without filtering on
a UHPLC separation with a strongly varying gradient baseline.
The Logarithmic Z in the View DAD/PDA 3D Matrix plots offers a third SymLog Z-scale option.
With the RdBu_r colormap, or another that goes to white at 0, you can judge the efficacy of the background
subtraction. Here we see the efficacy of the lsrpls at three different lambdas and without filtering on
a UHPLC separation with a strongly varying gradient baseline.