PeakLab v2 Documentation Contents R2N Software Home R2N Software Support
Import Spectroscopic Data Sets (DAD/PDA)
Import Spectroscopic Data Sets
This ChromSpec menu option is especially important for separations where the overlap in the fitted peaks in a chromatographic data set is incomplete and you can determine, from the fitting, that a single component exists between a given t1 and t2. You must first import spectra for this procedure to be available. This option allows you to extract the UV-VIS spectra for the pure component that was eluted during that time interval.

In this example, four distinct but chemically similar compounds are specified by their chromatographically exclusive time intervals in the separation:
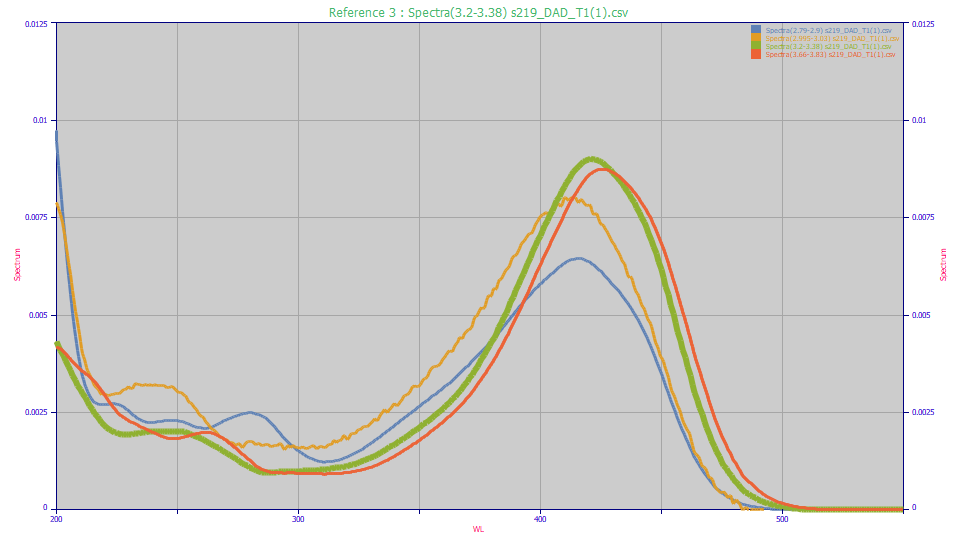
These UV spectra can then be fitted in PeakLab using Gaussian or Voigt functions in the Spectroscopy family. For comparison purposes, here we use the Unit Area option in the View and Compare Data option to see the four distinct UV and VIS principal absorptions. Note that low concentration peaks near the baseline (such as the amber curve) will have noisier spectra and those may require the Smooth option to get a clear picture of the peak wavelengths.
If you are using the first derivative method to remove one component of two coeluting peaks, you can use this approach with pure standards to get the UV-VIS maxima and minima wavelengths of each without running a separate analysis with a dedicated UV-VIS spectrometer. By using the chromatographic instrument's DAD to determine the exact wavelengths, these should be precisely the wavelengths needed in chromatographic analysis.